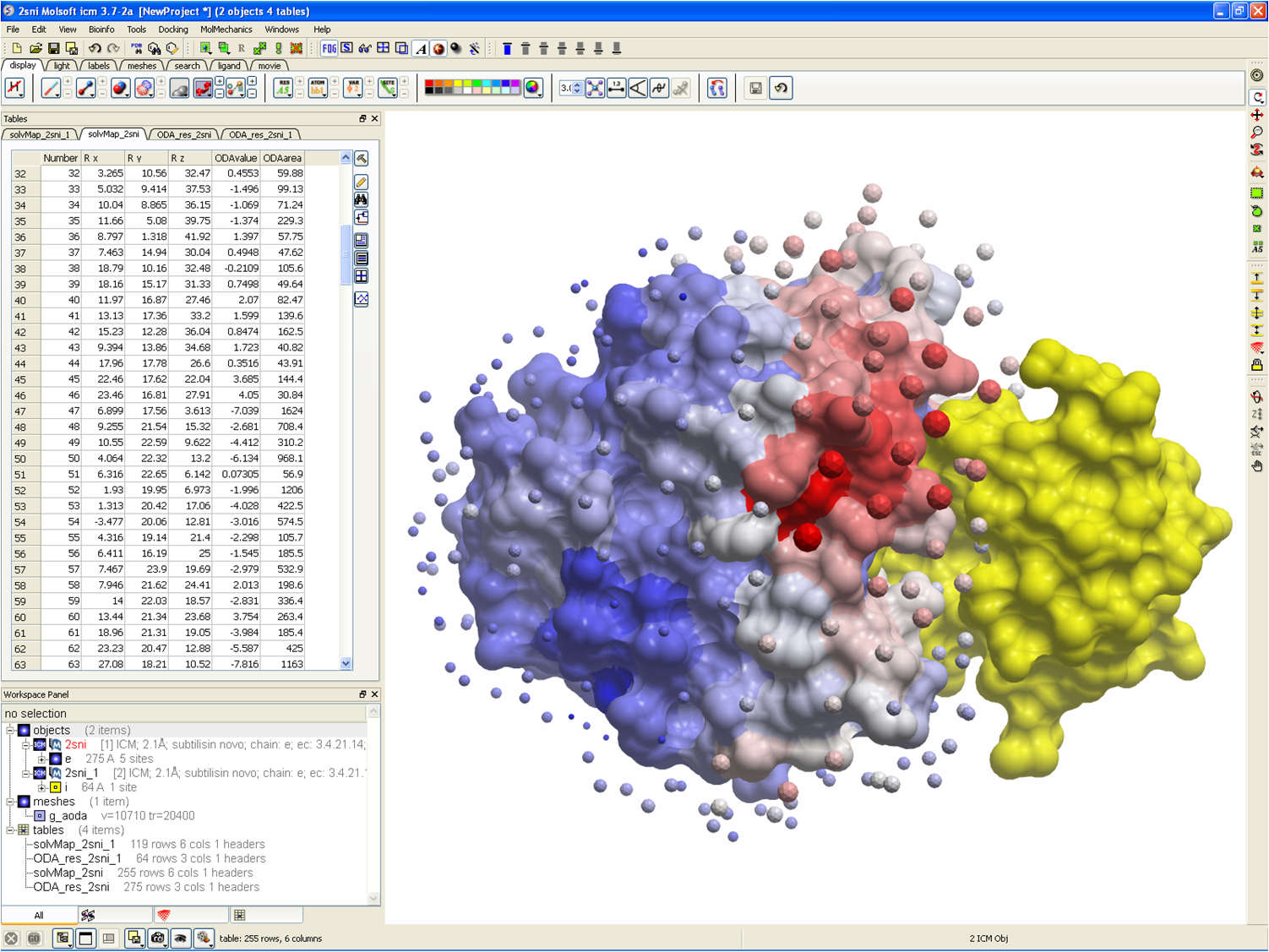
Protein-based Virtual Screening Tools applied for RNA-Ligand Docking identify new Binders of the preQ1-Riboswitch | bioRxiv
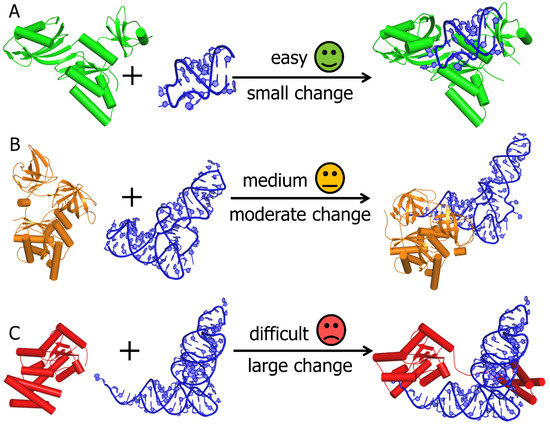
Genes | Free Full-Text | Bioinformatics Tools and Benchmarks for Computational Docking and 3D Structure Prediction of RNA-Protein Complexes
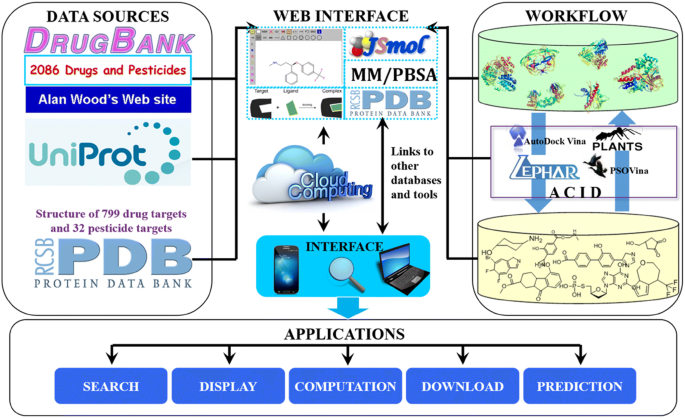
ACID: a free tool for drug repurposing using consensus inverse docking strategy | Journal of Cheminformatics | Full Text

Modelling peptide–protein complexes: docking, simulations and machine learning | QRB Discovery | Cambridge Core

MILCDock: Machine Learning Enhanced Consensus Docking for Virtual Screening in Drug Discovery | Journal of Chemical Information and Modeling

Protein-Based Virtual Screening Tools Applied for RNA–Ligand Docking Identify New Binders of the preQ1-Riboswitch | Journal of Chemical Information and Modeling

Virtual screening as a tool to discover new β-haematin inhibitors with activity against malaria parasites | Scientific Reports